One hundred seventy-six new packages made it to CRAN in July. Here are my “Top 40” picks organized into twelve categories: Data, Data Science, Finance, Genomics, Machine Learning, Mathematics, Medicine, Statistics, Time Series, Topological Data Analysis, Utilities and Visualization.
Data
eia v0.3.2: Provides API access to data from the US Energy Information Administration (EIA). Use of the API requires a free API key. See the Package Overview.
litteR v0.4.1: Implements a user interface to analyze litter data: beach litter, riverain litter, floating litter, seafloor litter, etc., in a consistent and reproducible manner. See the vignette for examples.
rSymbiota v1.0.0: Implements an interface to the Symbiota portals, allowing users to query taxon natural history collections that include plants, animals, and fungi. See the vignette to get started.
Data Science
bdpar v1.0.0: Provide a tool to easily build customized data flows to pre-process large volumes of information from different sources.
modeLLtest v1.0.0: Implements the cross-validated difference in means (CVDM) test by Desmarais and Harden (2014) and the cross-validated median fit (CVMF) test by Desmarais and Harden (2012). See the vignette to get started.
Finance
lazytrade v0.3.4: Provide sets of functions and methods to learn and practice data science using the idea of algorithmic trading. See the vignette.
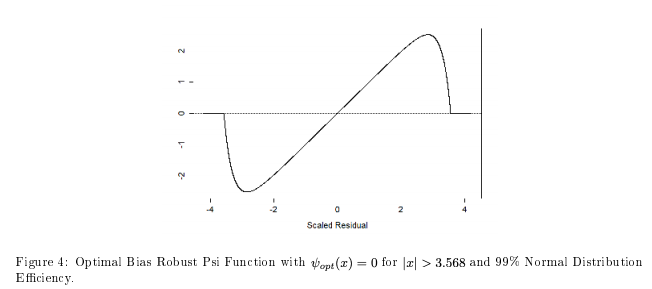
RPEIF v1.0: Computes the influence functions time series of the returns for the risk and performance measures as mentioned in Zhang and Martin (2017) as well as Chen and Martin (2018). The vignette provides some theory and examples.
Genomics
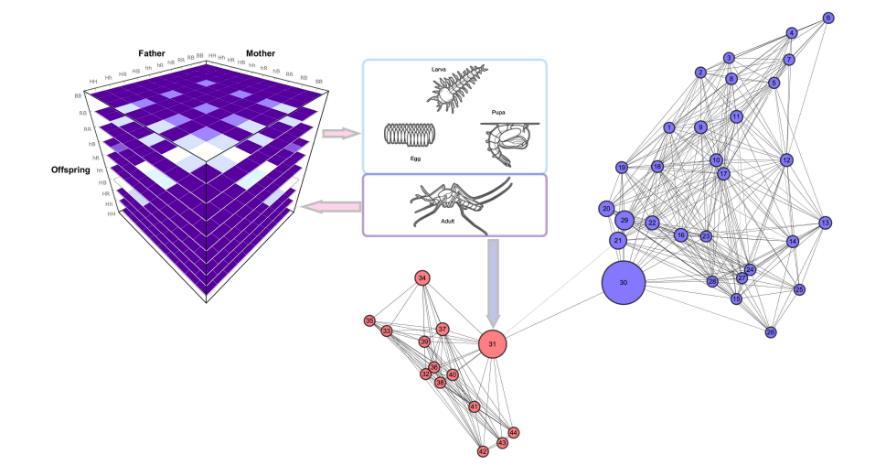
MGDrivE v1.0.0: Implements a testbed for gene drive interventions for mosquito-borne diseases control. See Deredec et al. (2001) and Hancock & Godfray (2007) for the background. There are vignettes for examples, the math, and a complete run.
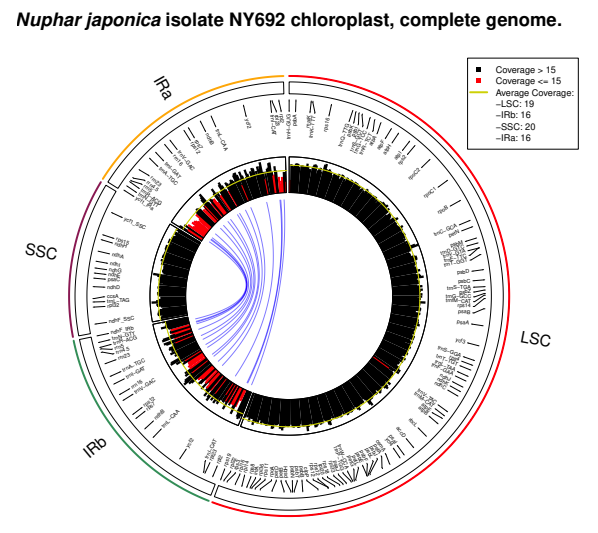
PACVr v0.8.1: Provides functions to visualize the coverage depth of a complete plastid genome, as well as the equality of its inverted repeat regions in relation to the circular, quadripartite genome structure and the location of individual genes. For details, see Gruenstaeudl and Jenke (2019), and for an example, see the vignette.
Machine Learning
forestRK v0.0-5: Provides functions that calculate common types of splitting criteria used in random forests for classification problems, as well as functions that make predictions based on a single tree or a Forest-R.K. The vignette offers an extended example.
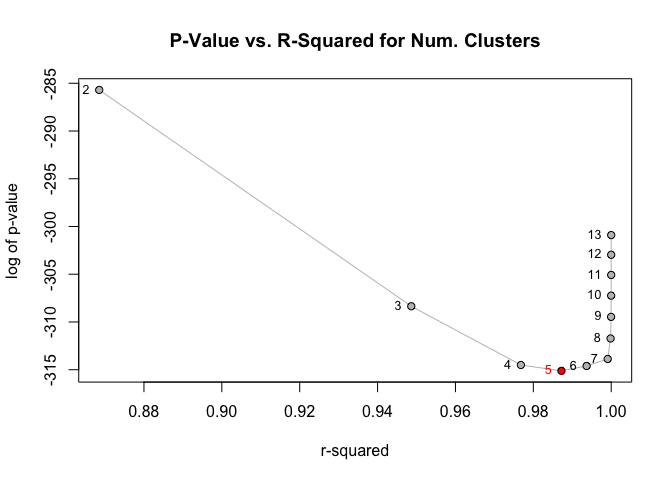
greenclust v1.0.0: Implements a method of iteratively collapsing the rows of a contingency table, two at a time, by selecting the pair of categories whose combination yields a new table with the smallest loss of chi-squared, as described by Greenacre (1988). See the README for an introduction to the package.
imgrec v0.1.0: Implements an interface to Vision AI, the Google image recognition system. See the vignette to get started.
mlr3 v0.1.2: Provides R6 object-oriented programming building blocks for machine learning tasks. Look here for a brief overview.
Mathematics
odin v1.0.1: Provides functions to generate systems of ordinary differential equations (ODE) and integrate them, using a domain specific language (DSL). The DSL uses R’s syntax, but compiles to C to efficiently solve the system. There is a vignette on Discrete Models and another with several examples.
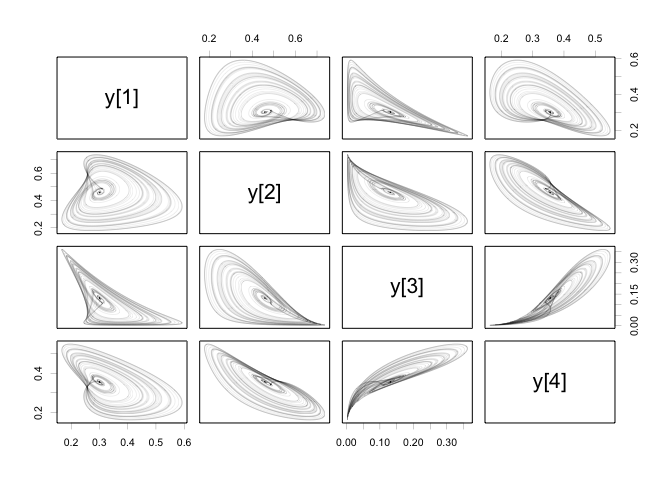
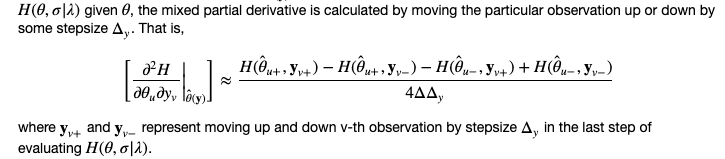
pCODE v0.9.2: Contains an implementation of the parameter cascade method from Ramsay, J. O., Hooker,G., Campbell, D., and Cao, J. (2007) for estimating ordinary differential equation models with missing or complete observations. The vignette shows the math.

Medicine
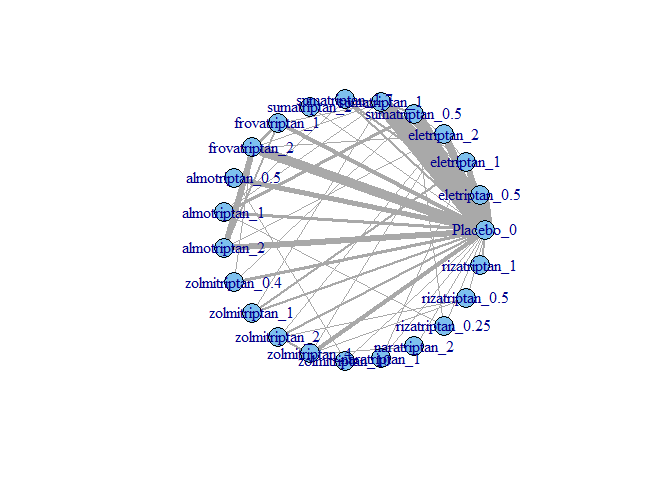
MBNMAdose v0.2.3: Provides functions to fit Bayesian dose-response, model-based network meta-analysis (MBNMA) that incorporate multiple doses within an agent by modelling different dose-response functions. See Mawdsley et al. (2016) for the theory and the vignette for examples.
qMRI v1.0.1: Implements methods to estimate quantitative maps described in Weiskopf et al. (2013). See the vignette for an example.
Statistics
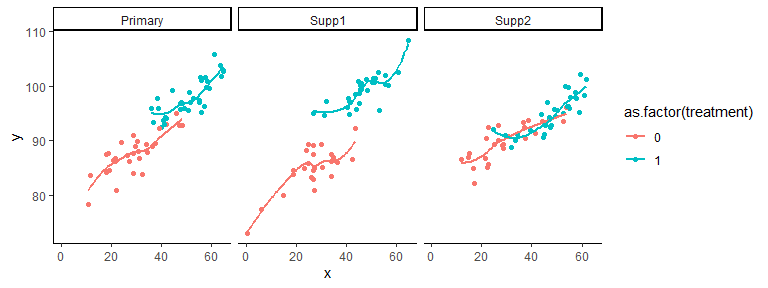
borrowr v0.1.0: Provides functions to estimate population average treatment effects from a primary data source with borrowing from supplemental sources. Causal estimation is done with either a Bayesian linear model or with Bayesian additive regression trees (BART) to adjust for confounding. See Chipman et al.(2010) Borrowing is done with multisource exchangeability models (MEMs). See Kaizer et al. (2018). There is a vignette.
emax.glm v0.1.2: Implements Expectation Maximization (EM) regression for general linear models. See Hastie et al. (2009) and Zeileis et al. (2017) for background. There is an introduction and vignettes on the EM GLM algorithm, Residual Theory and on Warm up and Exposure in the GLM algorithm.
kosel v0.0.1: Implements functions to perform variable selection for many types of L1-regularised regressions using the revisited knockoffs procedure. See Gegout et al. (2019) for the details.
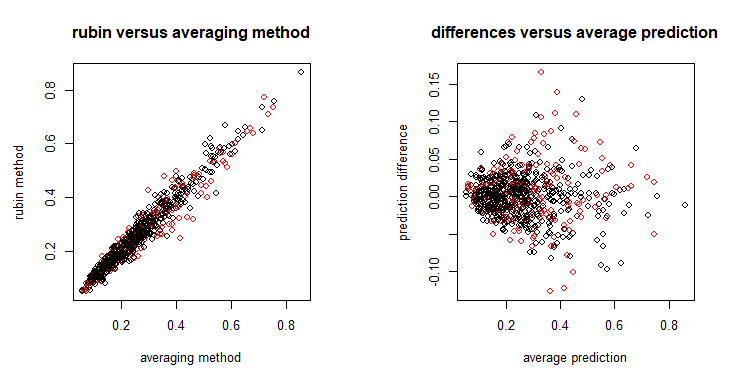
mipred v0.0.1: Calibrates generalized linear models and Cox regression models for prediction using multiple imputation to account for missing values in the predictors. See Mertens et al. (2018) for the method and the vignette for examples.
MixMatrix v0.2.2: Provides sampling and density functions for matrix variate normal, t, and inverted t distributions using the EM algorithm. See Thompson et al. (2019) for background. There are vignettes on Discriminant Analysis, matrix-t-estimation, the Matrix Normal Distributions, and Mixture Models.
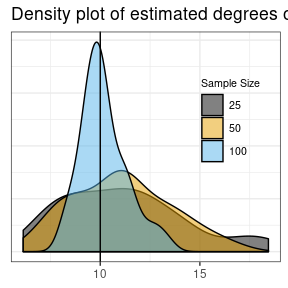
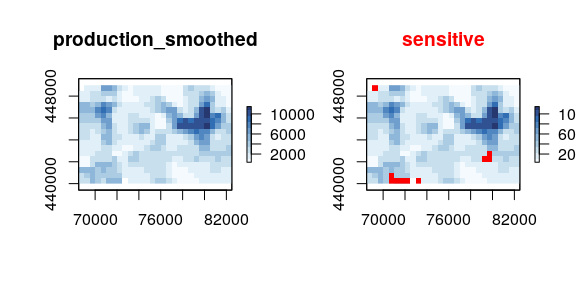
sdcSpatial v0.1.1: Provides functions to create privacy protected raster maps can be created from spatial point data. Protection methods include smoothing of dichotomous variables by de Jonge and de Wolf (2016), continuous variables by de Wolf and de Jonge (2018), suppressing revealing values and a generalization of the quad tree method by Suñé et al. (2017).
Time Series
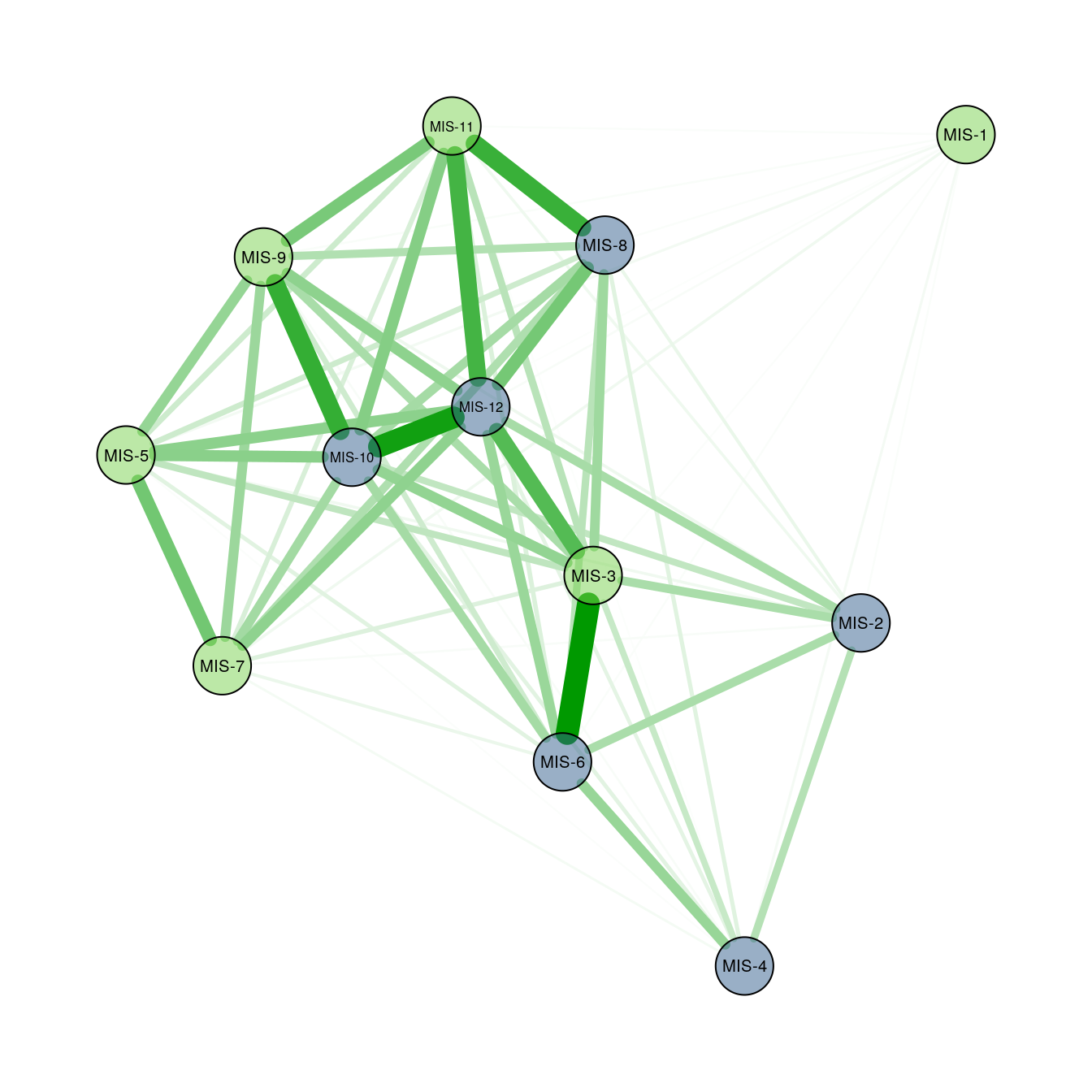
distanta v1.0.1: Provides tools to assess the dissimilarity between multivariate time-series. It is based on the psi measure described by Birks and Gordon (1985), which computes dissimilarity between irregular time-series constrained by sample order. There is a Package Summary with examples.
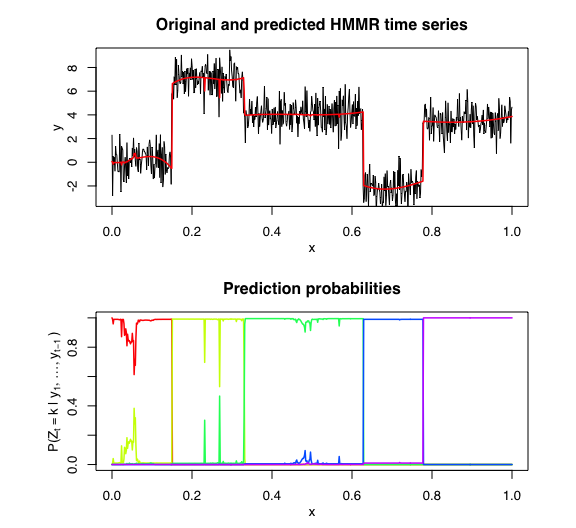
samurais v0.1.0: Provides a variety of statistical latent variable models and unsupervised learning algorithms to segment and represent both univariate and multivariate time-series data. There are vignettes for (1) Hidden Markov Model Regression (HMMR), (2) Model Selection for HMMR, (3) Multivariate HMMR, (4) Model Selection for MHMMR, (5) Regression with Hidden Logistic Process (RHLP), (6) [Model selection for RHLP, (7) Multivariate RHLP, (8) Model Selection for MRHLP, and (9) Piecewise Regression.
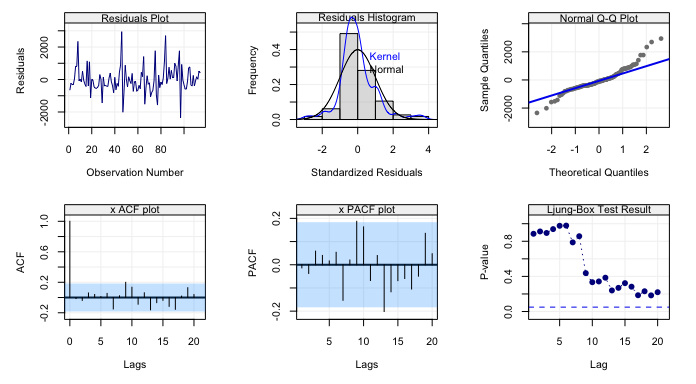
simts v0.1.1: Implements a system of tools to support time series analysis courses, including a technique called Generalized Method of Wavelet Moments (GMWM). See Guerrier et al. (2013) for the background and the vignette for examples.
Topological Data Analysis
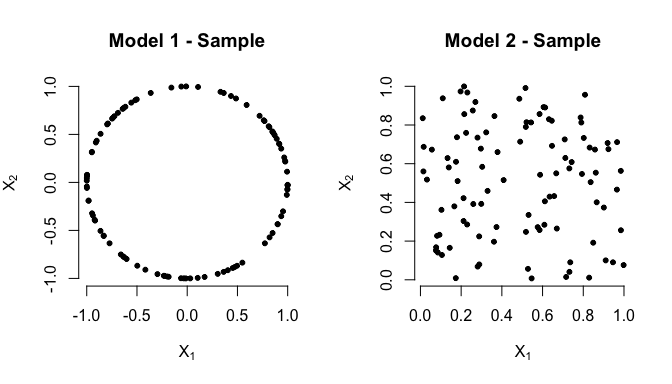
BallMapper v0.2.0: Provides functions to compute a topologically accurate summary of data in the form of an abstract graph, using the algorithm described in Dlotko, (2019). Have a look at this video for an overview.
kernelTDA v0.1.1: Provides tools for exploiting topological information in standard statistical learning algorithms, implements kernels defined on the space of persistence diagrams, and provides a solver for kernel Support Vector Machines based on the C++ LIBSVM and functions to compute Wasserstein distance using the C++ HERA library. The vignette offers examples.
Utilities
babelwhale v1.0.0: Provides a unified interface to interact with docker and singularity containers, allowing users to execute a command inside a container, mount a volume, or copy a file.
fastmap v1.0.0: Provides a fast implementation of a key-value store that avoids common memory leakage issues by using data structures in C++. See the README for an overview.
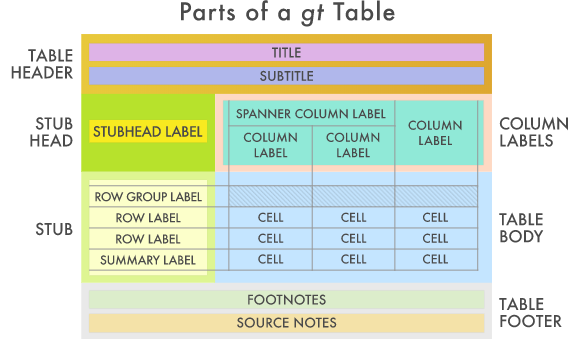
modelsummary v0.1.0; Leverages the gt and broom packages to create customizable, publication-ready summary tables for statistical models. See the GitHup Repo for an overview of how to use the package.
readwritesqlite v0.0.2: Provides functions to read and write data frames to SQLite databases while preserving time zones (for POSIXct columns), projections (for sfc columns), units (for units columns), levels (for factors and ordered factors), and classes for logical, Date, and hms columns. The vignette provides usage information.
rolldown v0.1: Provides R Markdown output formats based on JavaScript libraries such as Scrollama for storytelling. There are vignettes on Basic Style and Sidebar Layout for Scrollama documents.
rray v0.1.0: Provides a toolkit for manipulating arrays in a consistent, powerful, and intuitive manner through the use of broadcasting and a new array class, rray. There are vignettes on Broadcasting, The rray, and Toolkit.
wyz.code.offensiveProgramming v1.1.8: Provides code to ease the transition from defensive to offensive programming as described in the Offensive Programming Book.
Visualization
altair v3.1.1: Implements an interface to Altair, which itself is a Python interface to Vega-Lite.

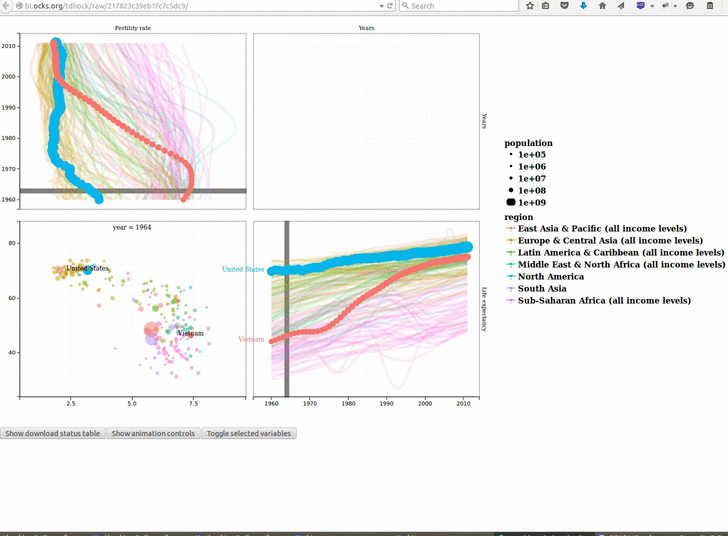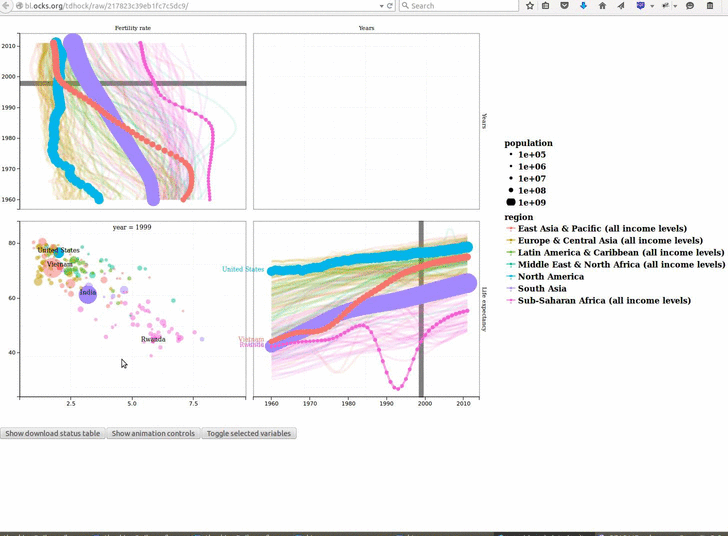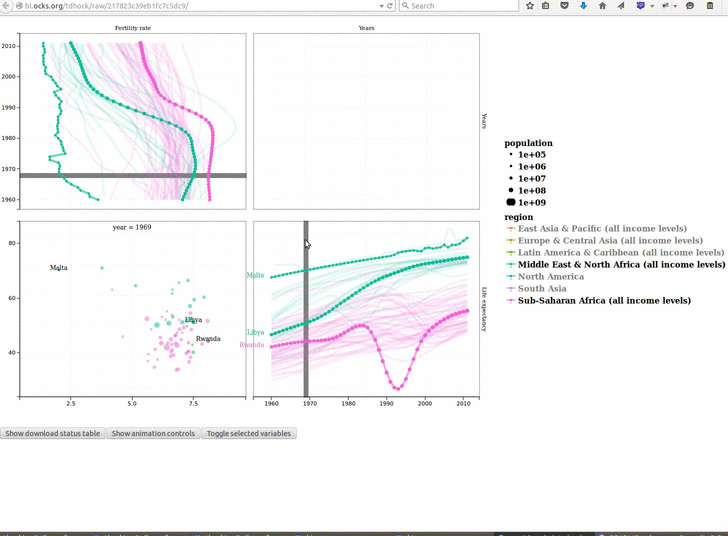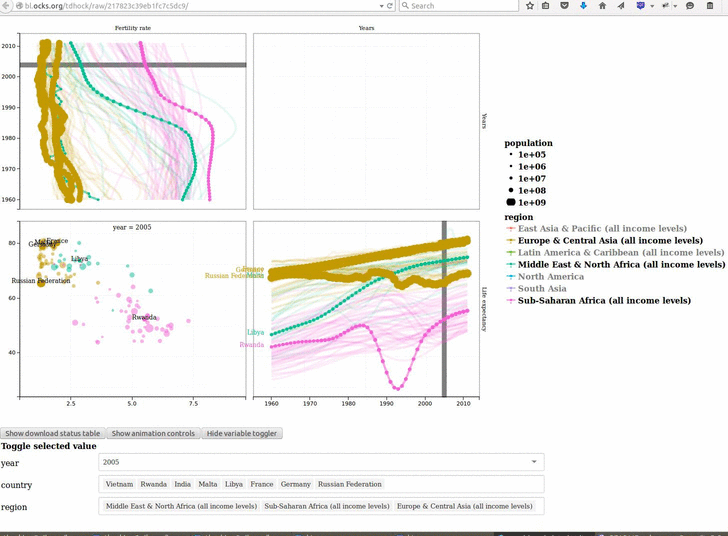
animint2 v2019.7.3: Provides functions for defining animated, interactive data visualizations in R code, and rendering on a web page. See the 2018 Journal of Computational and Graphical Statistics paper for a description of the underlying concepts, and the Github page for examples.
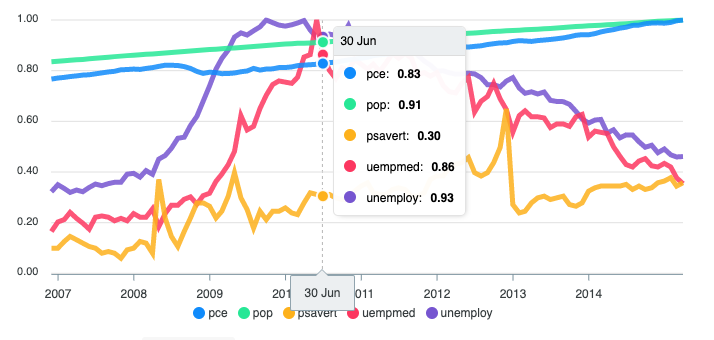
apexcharter v0.1.2: Provides an htmlwidgets interface to apexcharts.js, a modern JavaScript charting library to build interactive charts and visualizations with simple API. There is a Getting Started Guide and vignettes on labs and lines.
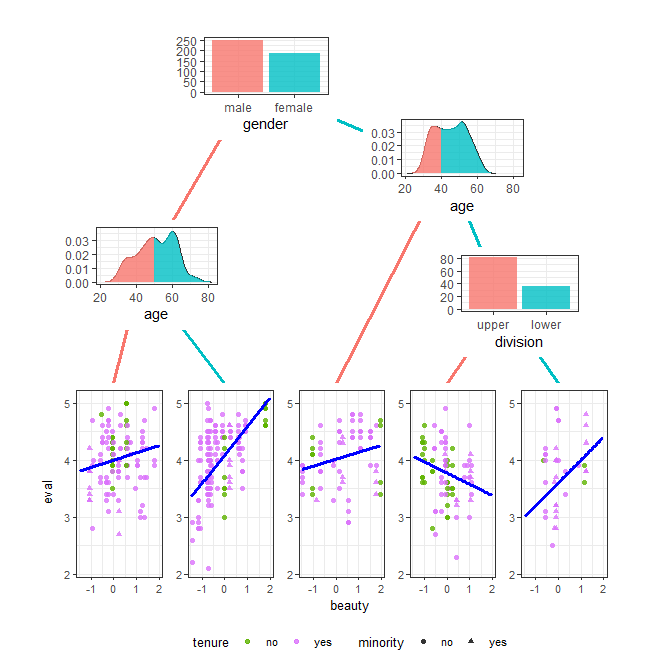
ggparty v1.0.0: Extends ggplot2 functionality to the partykit package, which provides the tools to create structured and highly customizable visualizations for tree-objects of the class party. There is a vignette on ggparty and another on edges.
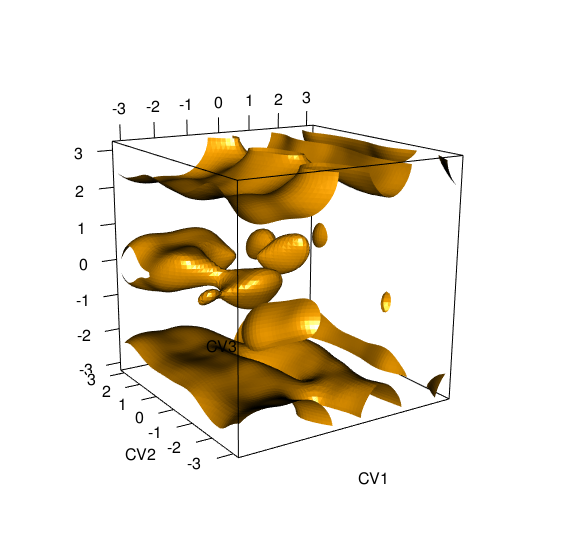
metadynminer3d v0.0.1: Provides tools to read, analyze, and visualize Metadynamics 3D HILLS files from Plumed. See Tribello et al. (2014) for the background and look here for examples.
You may leave a comment below or discuss the post in the forum community.rstudio.com.