One hundred sixteen new packages stuck to CRAN in December 2022. Here are my “Top 40” selections in thirteen categories: Computational Methods, Data, Ecology, Epidemiology, Genomics, Machine Learning, Mathematics, Medicine, Networks, Signal Processing, Statistics, Utilities, and Visualization.
Computational Methods
bgw v0.1.0: Implements the BGW algorithm described by Bunch, Gay and Welsch (1993) which exploits the special structure of statistical estimation problems within a trust-region-based optimization approach to produce an estimation algorithm that is much more effective than the usual practice of using optimization methods originally developed for general optimization. See the vignette.
GPArotateDF v2022.12-1: Provides functions for derivative-free gradient projection algorithms for factor rotation. See Jennrich (2004) and Bernaards and Jennrich (2005) for the theory, and the vignette for examples.
Mestim v0.2.1: Implements a flexible framework for estimating the variance-covariance matrix of parameters by computing the empirical sandwich variance. See Tsiatis et al. (2019) for background and the vignette for some theory and examples.
Data
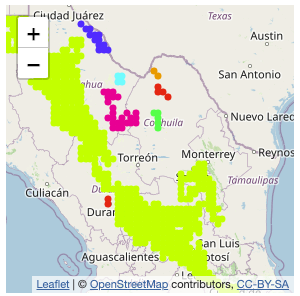
dafishr v1.0.0: Provides functions to download, clean and analyse raw Vessel Monitoring System, VMS, data from Mexican government. See the vignette to get started.
hcidata v0.1.0: Provides a collection of datasets of human-computer interaction (HCI) experiments. Each dataset is from an HCI paper, with all fields described and the original publication linked. The datasets sources include Bergström et al. (2022), Dalsgaard et al. (2021), Larsen et al. (2019), Lilija et al. (2019), Pohl and Murray-Smith (2013), and Pohl and Mottelson (2022).
Ecology
bamm v0.4.3: Implements species distribution models as a function of biotic and movement factors. See Soberón and Osorio-Olvera (2022) for the theoretical background and the vignette for examples.
gadget3 v0.8-4: Provides a framework for creating marine ecosystem models that generatesR or C++ code which can then be optimized using the TMB package and standard tools. See Kristensen et al. (2016) and Begley & Howell (2004) for background and the vignettes Model Debugging, Model Structure, and Writing G3 Actions.
Epidemiology
finalsize v0.1: Provides functions to calculate the final size of a susceptible-infectious-recovered epidemic in a population with demographic variation in contact patterns and susceptibility to disease. See Miller (2012) for details. Additionally, there is an Introduction and vignettes on modeling uncertainty in R0 and modeling heterogeneous susceptibility.

Genomics
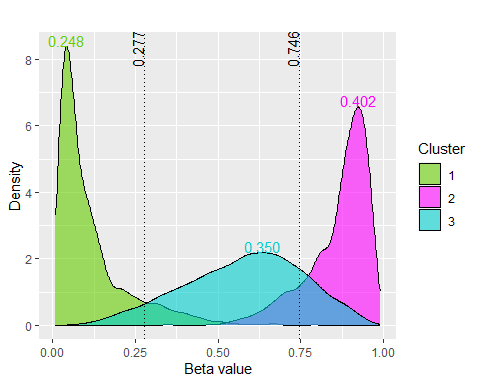
betaclust v1.0.0: Implements the family of novel beta mixture models developed by Majumdar et al. (2022) to appositely model the beta-valued cytosine-guanine dinucleotide (CpG) sites, to objectively identify methylation state thresholds, and to identify the differentially methylated CpG (DMC) sites using a model-based clustering approach. See the vignette.
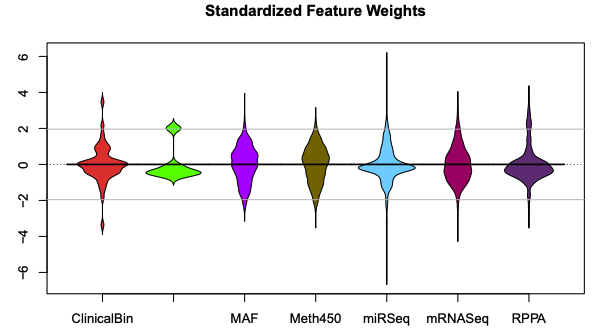
plasma v0.9.21: Implements tools for supervised analyses of incomplete, overlapping multiomics datasets. Functions apply partial least squares in multiple steps to find models that predict survival outcomes. See the vignettes on Interpretation and Partial Least Squares.
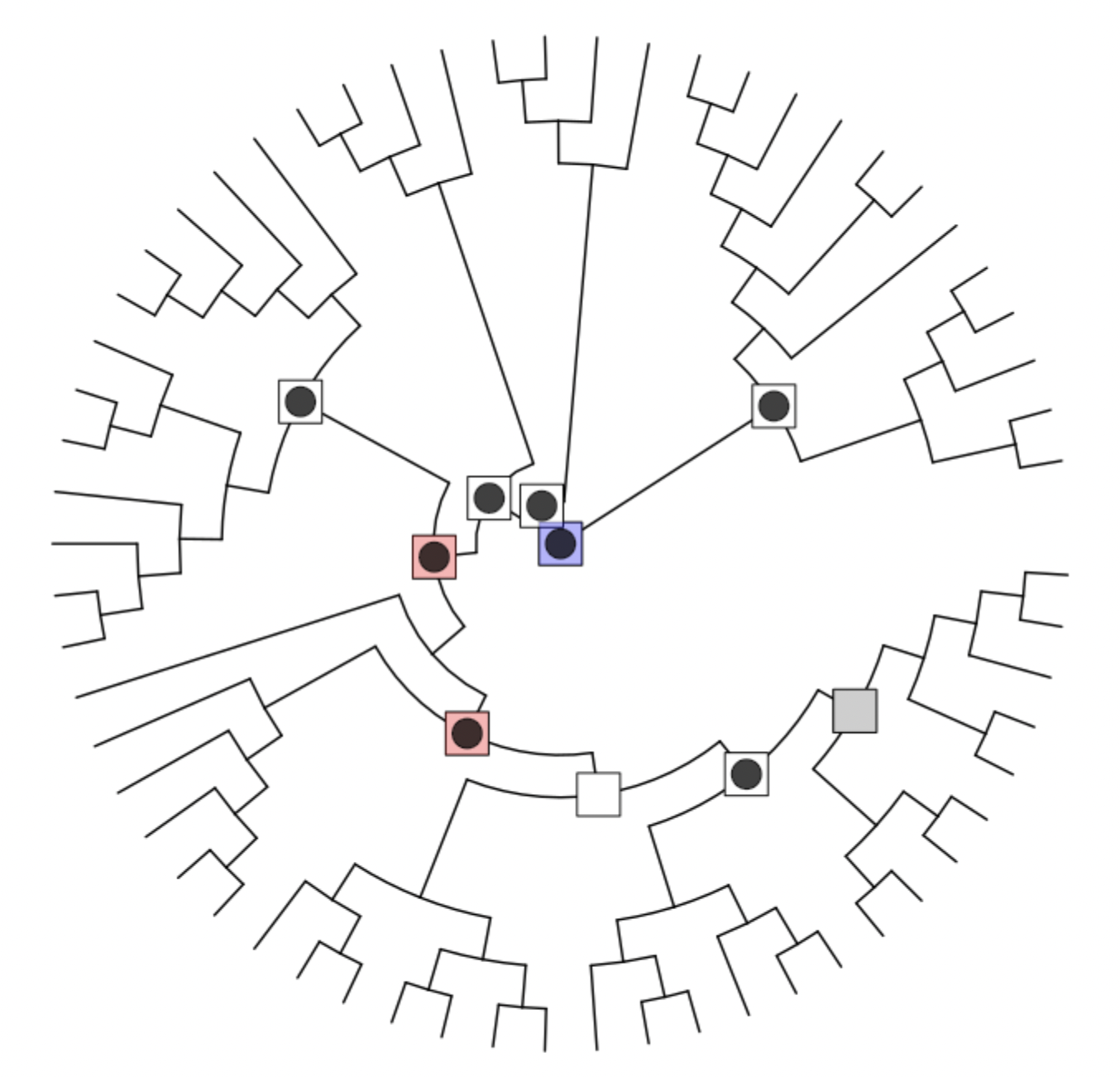
POMS v1.0.1: Provides code to identify functional enrichments across diverse taxa in phylogenetic tree, particularly where these taxa differ in abundance across samples in a non-random pattern. See Douglas et al. (2022) for background and look here for documentation.
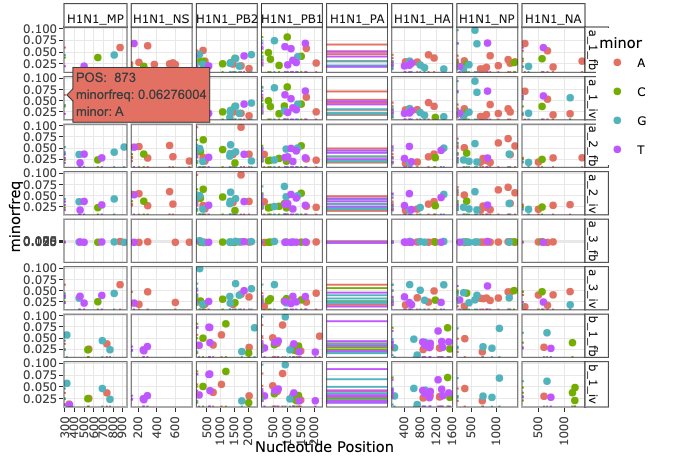
vivaldi v1.0.0: Provides functions to analyze minor alleles of viral genomes from sequencing data of viral genomes in Illumina vcf files. See the vignette.
Machine Learning
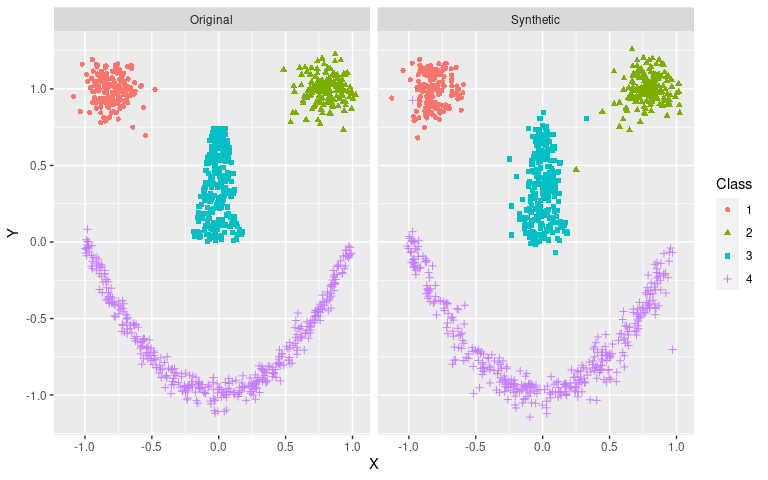
arf v0.1.2: Implements adversarial random forests, an iterative algorithm that recursively partition data into fully factorized leaves, where features are jointly independent. See Watson et al. (2022) for details and the vignette on Density Estimation.
correctR Provides corrected test statistics for cases when samples are not independent, such as when classification accuracy values are obtained over resamples or through k-fold cross-validation, as proposed by Nadeau and Bengio (2003) and presented in Bouckaert and Frank (2004). See the vignette.
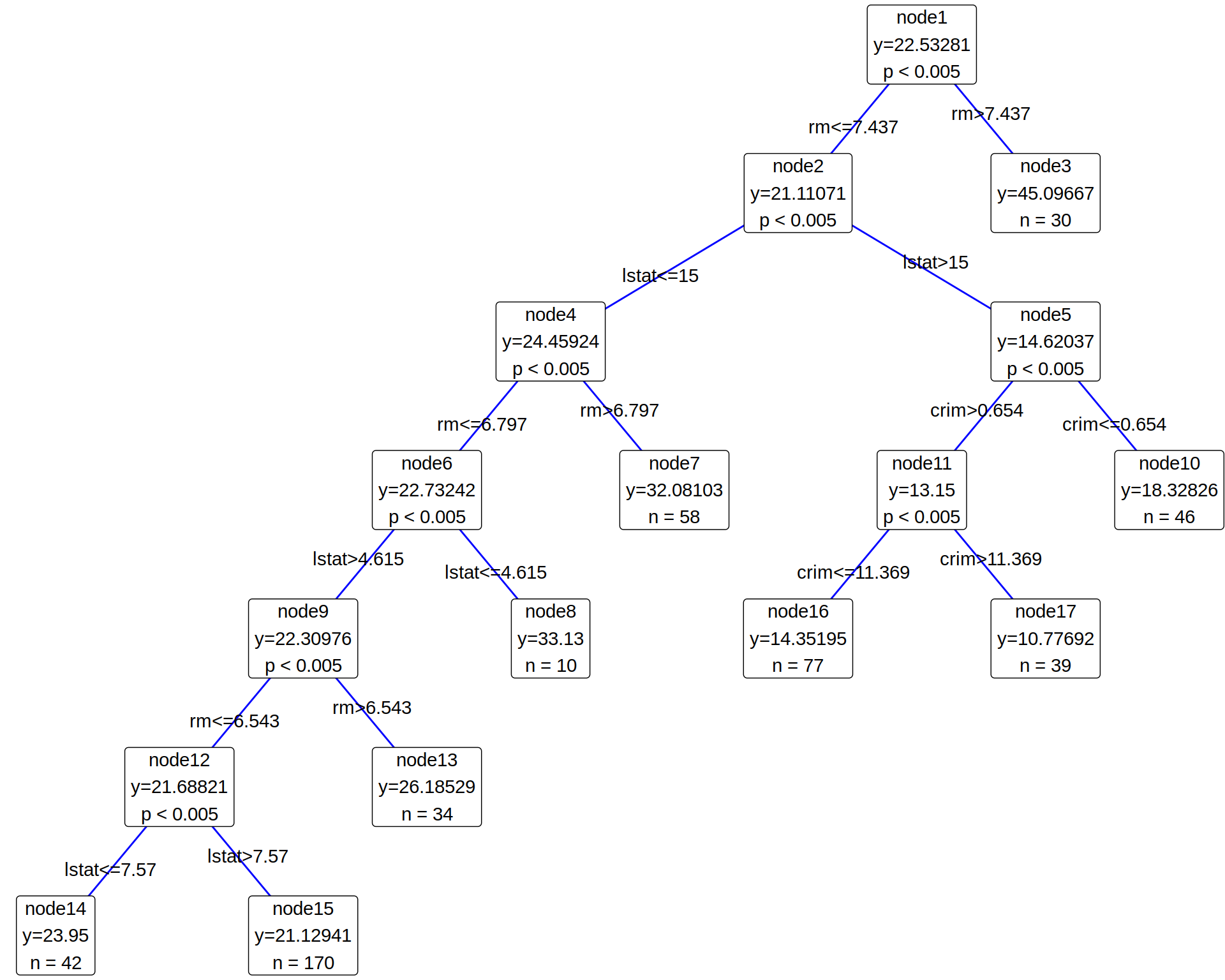
HTT v0.1.1: Implements a novel decision tree algorithm in the hypothesis testing framework that examines the distribution difference between two child nodes over all possible binary partitions. The test statistic enables the algorithm to better detect complex structure and not only the mean difference. See the vignette
janus v1.0.0: Implements a tensorflowbased, deep-neural network recommending system with coarse-to-fine optimization. Look here for an introduction.
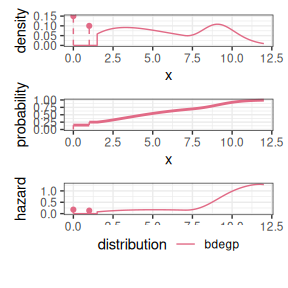
reservr v0.0.1: Provides functions to fit distributions and neural networks to censored and truncated data. See Bücher & Rosenstock (2022) for background and the vignettes Working with Distributions and TensorFlow Integration.
Mathematics
qspray v0.1.1: Provides functions for the symbolic calculation and evaluation of multivariate polynomials with rational coefficients. See README for an example.
Medicine
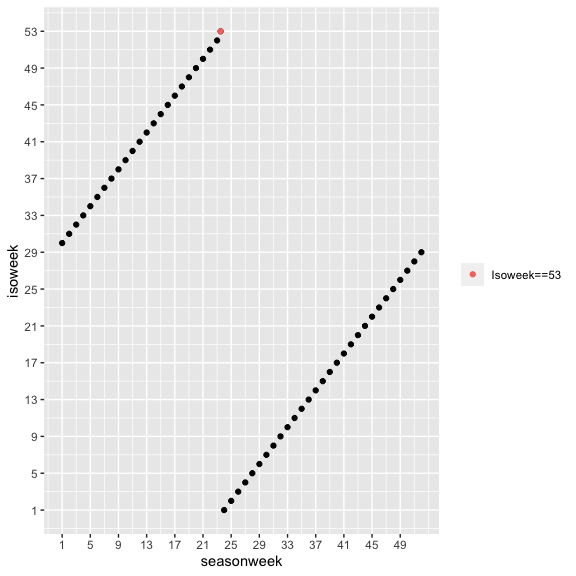
cstime v2022.11.22: Provides consistent time conversion functions for public health purposes including conversions between date, ISO week, ISO yearweek, ISO year, calendar month/year, season, and season week. There is an Introduction and there are additional vignettes Date, year, week conversion and Season week.
GNGTools v1.0.0: Implements a Go/No-Go decision-making framework based on Bayesian posterior probabilities linked to the target product profile. See the vignette.
oncomsm v0.1.2: Implements methods to fit a parametric Bayesian multi-state model to tumor response data. The model can be used to sample from the predictive distribution, to impute missing data, and to calculate probability of success for custom decision criteria during an ongoing trial. See the vignettes avoiding-bias, oncomsm, and prior-choice.

REDCapDM v0.1.0: Provides functions to to read and manage REDCap data and identify missing or extreme values. See the vignette.
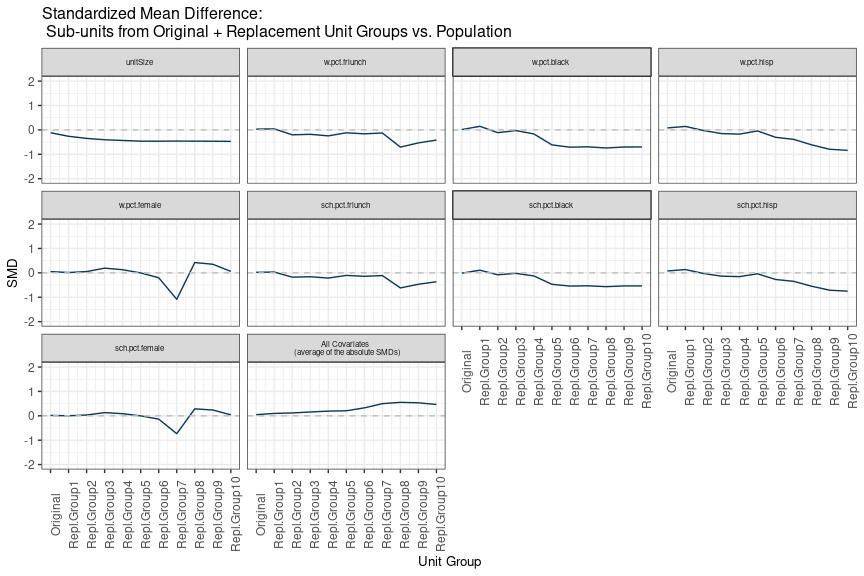
sitePickR v0.0.1: Provides functions to perform a two-level process to select a representative sample of sites for a prospective study such as a randomized controlled trial where the possibility of an initially selected site may not want to participate is anticipated. See Deville & Tillé (2004) and Tillé (2011) for background on the method employed and the vignette for a demo.
Networks
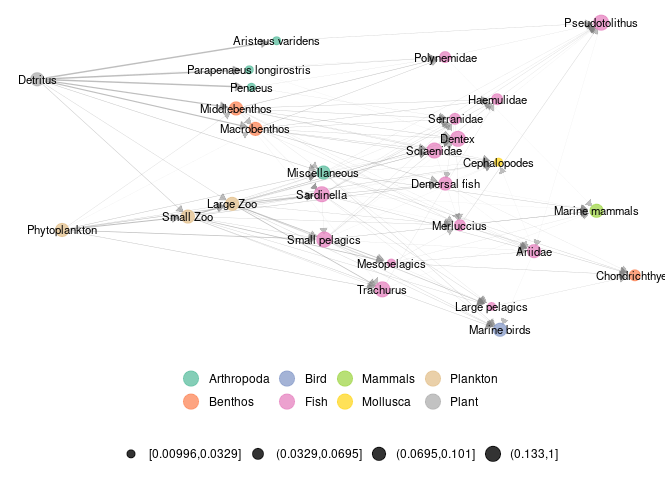
metanetwork v0.7.0: Provides a toolbox to represent large trophic networks in space or time across aggregation levels. The layout algorithm uses dimension reduction on a diffusion graph kernel and trophic levels. Look here for examples.
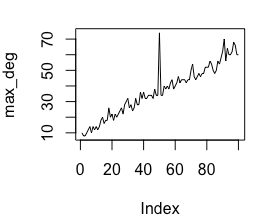
oddnet v0.1.0: Implements a feature-based method to identify anomalies in dynamic, temporal networks. See Kandanaarachchi & Hyndman (2022) for the theory and the vignette for an example.
Signal Processing
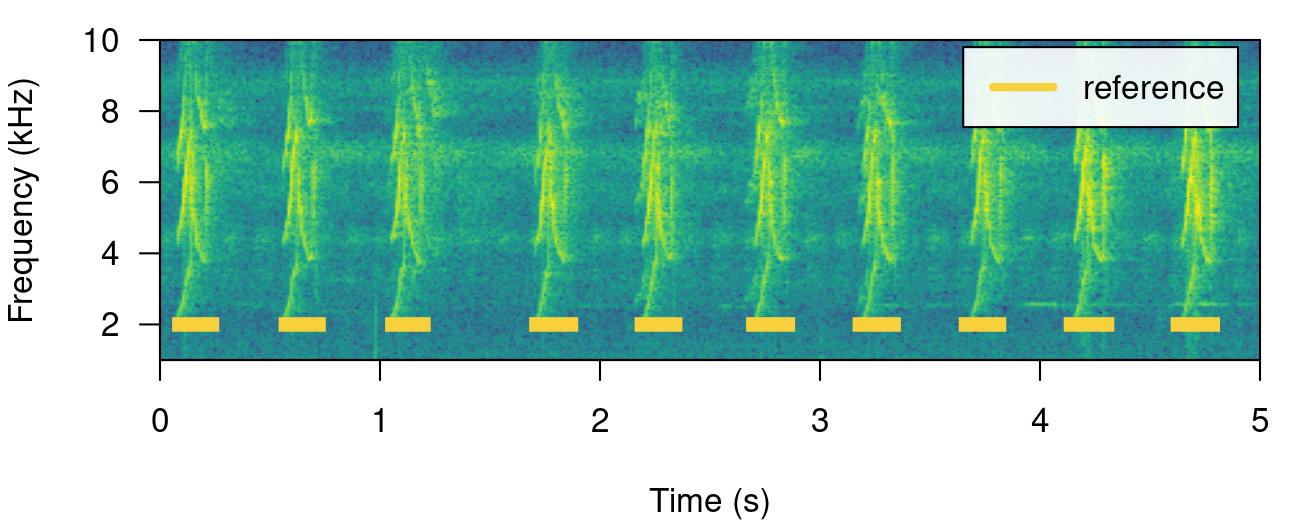
ohun v0.1.0: Provides functions to facilitates the automatic detection of acoustic signals, including functions to diagnose and optimize the performance of detection routines. See Hossin & Sulaiman (2015) for background and the vignette for examples.
Statistics
gmvjoint v0.1.0: Implements an EM algorithm for fitting joint survival and glm models of longitudinal data. See Bernhardt (2015) for background and README for examples.
hdcate v0.1.0: Implements a two-step double-robust method to estimate the conditional average treatment effects (CATE) with potentially high-dimensional covariates as described in Fan et al. (2022). See the vignette.
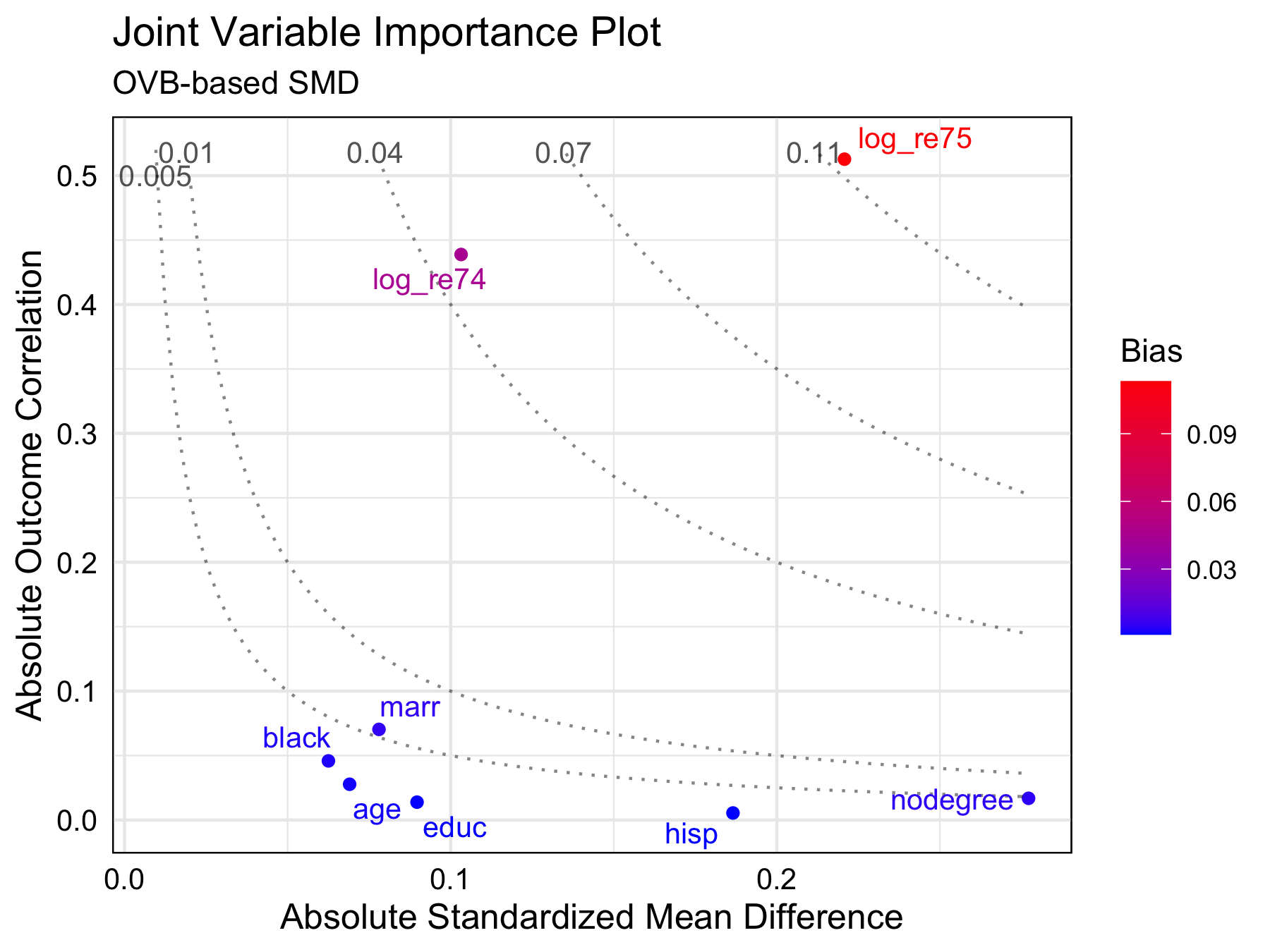
jointVIP v0.1.1: Implements joint variance importance plots to assist with variable selection and parameter tuning. See by Liao et al. (2023) for background. There is a Getting Started Guide and a vignette on Options.
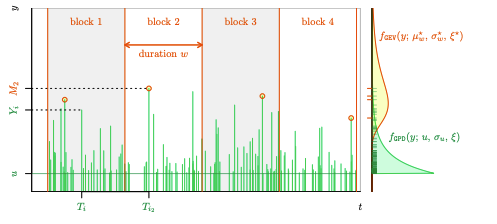
nieve v0.1.1: Provides utility functions and objects for extreme value analysis including probability functions with their exact derivatives and transformations exchanging the two parameterizations of peaks over threshold models. See the vignette.
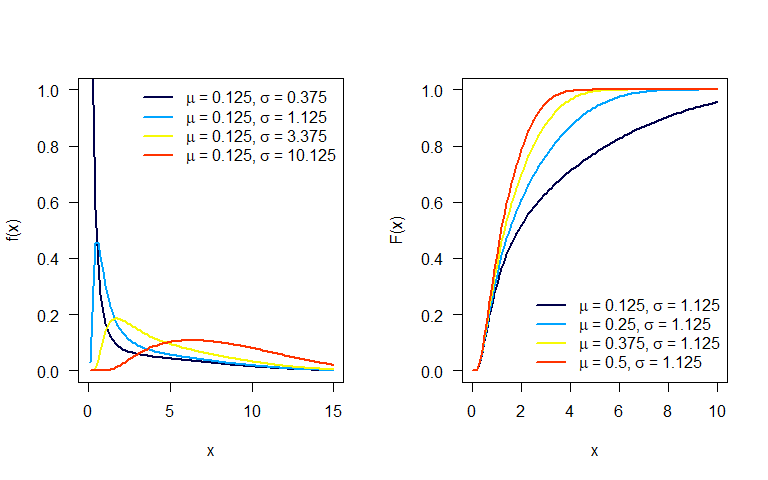
RelDists v1.0.0: Provides functions for parameter estimation and linear regression models for the reliability distribution families reviewed by Almalki & Nadarajah (2014). Generalized Additive Models are used for location, scale and shape. See Rigby & Stasinopoulos (2005). There are vignettes on the [FWE]() and OQ distributions.
ridgetorus v1.0.1: Implements Principal Component Analysis (PCA) on the torus via density ridge estimation, and includes functions for evaluating, fitting, and sampling these models. See García-Portugués and Prieto-Tirado (2022) for the theory and the vignette for examples.
Utilities
dataset v0.2.0: Implements a subjective interpretation of the W3C DataSet recommendation and the datacube model, which relevant to the global Statistical Data and Metadata eXchange standards, the Dublin Core, and the DataCite standards preferred by European open science repositories to improve the findability, accessibility, interoperability and reusability of the datasets. There are several vignettes including Motivation and Datasets with FAIR Metadata.
nlmixr2rpt v0.1.0: provides functions to generate reporting workflows around nlmixr2 analyses with outputs in Word and PowerPoint including functions to specify figures, tables and report structure in a user-definable YAML file. See the vignettes Accessing Figures and Tables and Reporting nlmixr2 Fit Results.
qtwAcademic v2022.12.13: Provides Quarto website templates commonly used by academics including templates for personal websites and course/workshop websites. There is an Intraduction and there are vignettes for workshop and personal websites.
r2social v1.0: Provides JavaScript and CSS styles to allow easy incorporation of various social media elements in shiny applications, dashboards and rmarkdown documents. The elements include share buttons, connect with us buttons, and hyperlink buttons. See the vignette.
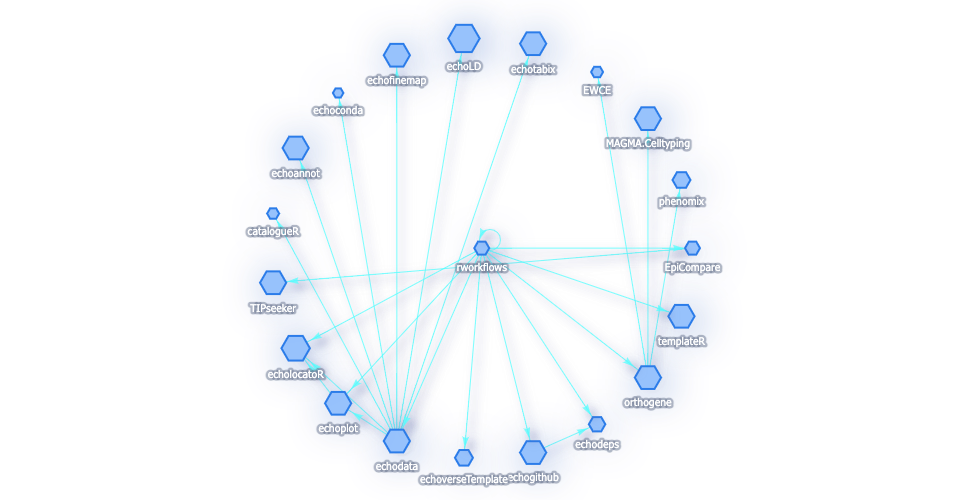
rworkflows v0.99.5: Implements functions for the continuous integration for R packages, automated testing, website building, and containerised deployment. See the vignettes depgraph, docker, repositories report, and rworlflows.
shinyDatetimePicker v1.0.0: Provides three types of datetime pickers for usage in a shiny UI. See README for examples.
Visualization
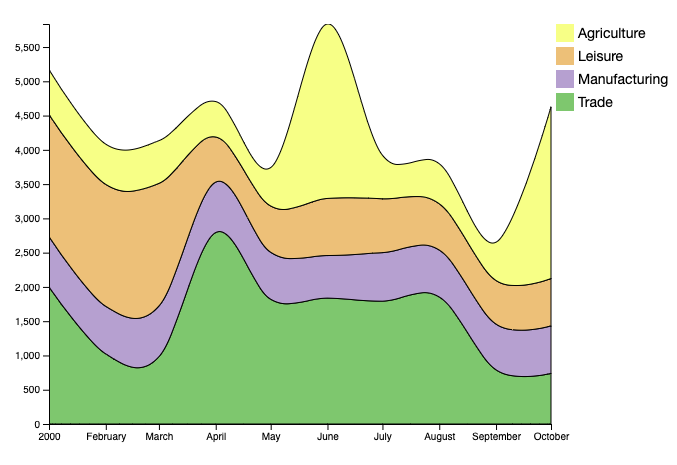
ddplot v0.0.1: Implements an API to allow users to create D3 based SVG plots using the r2d3 library. See the vignette.
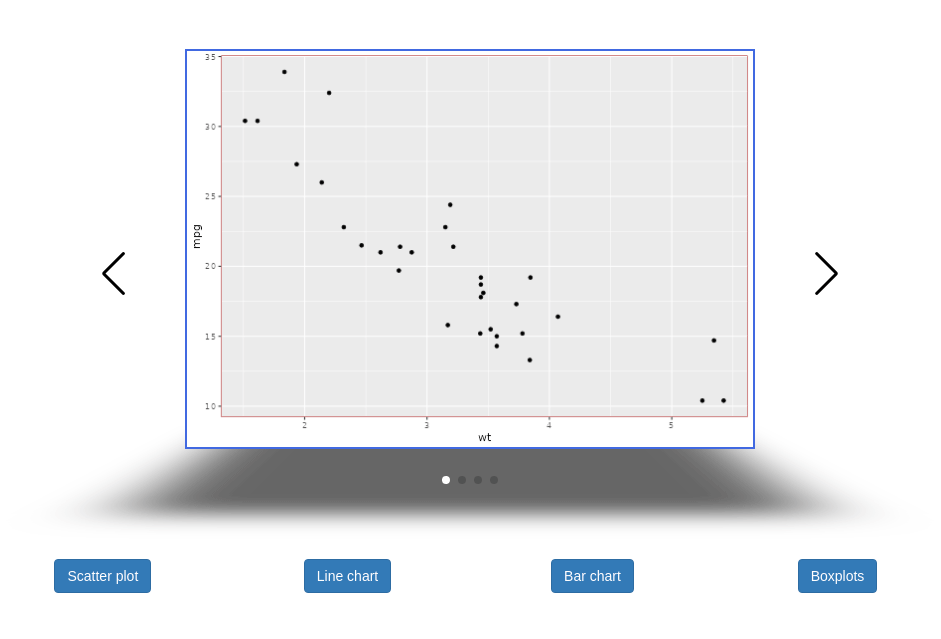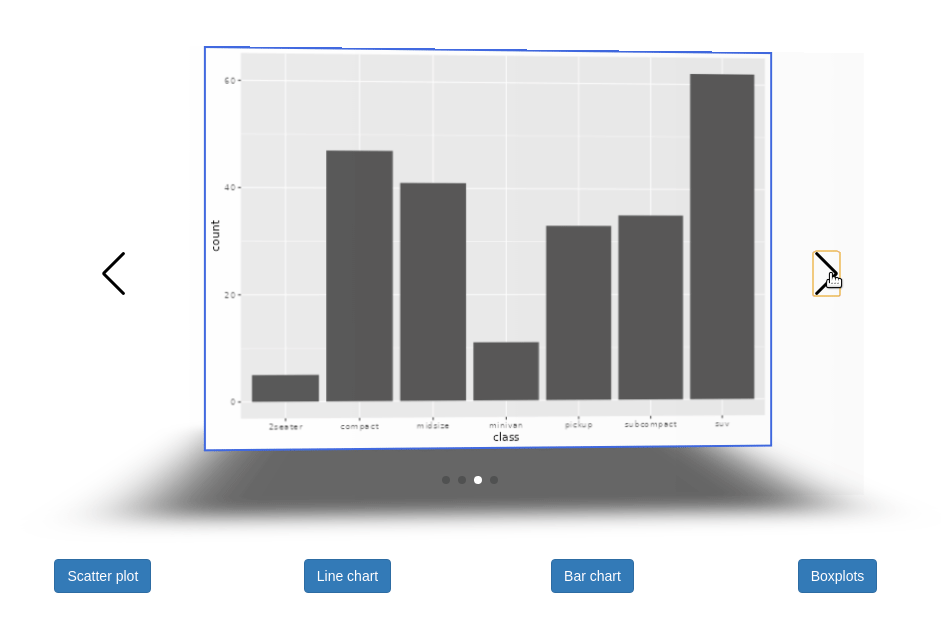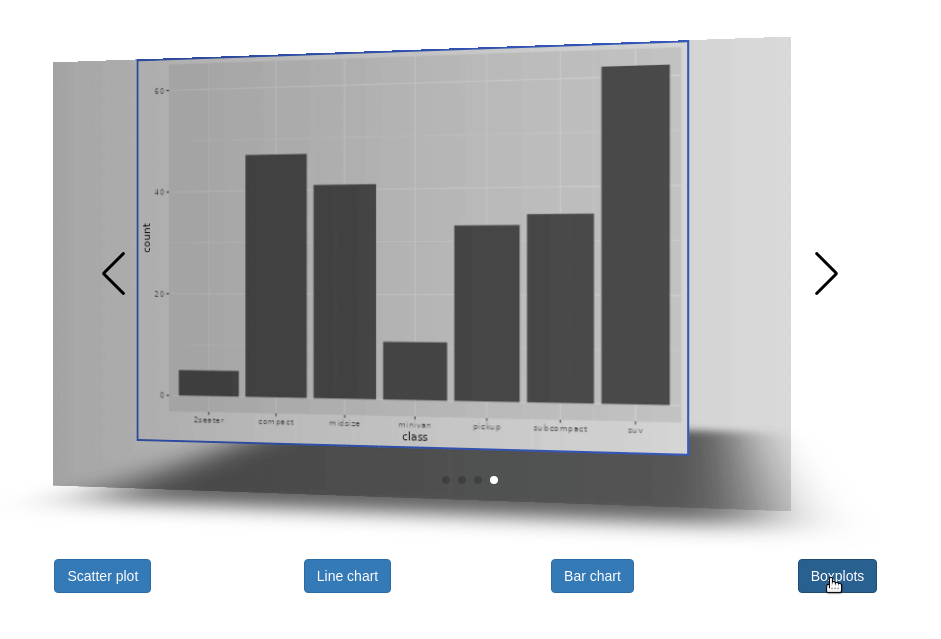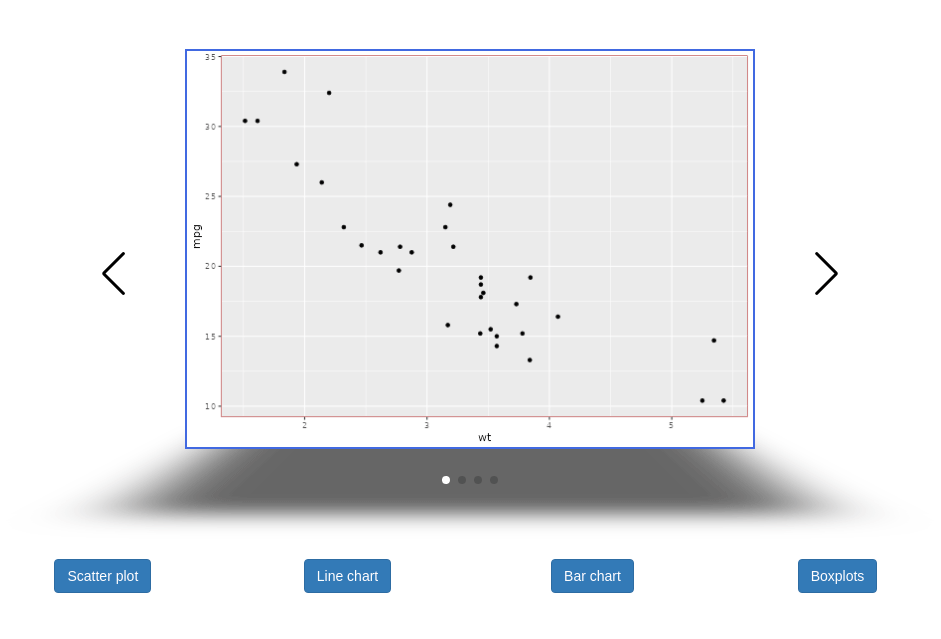
swipeR v0.1.0: Provides tools to create carousels using the JavaScript library Swiper and htmlwidgets. Carousels can be displayed in the RStudio viewer pane, in Shiny applications and in rmarkdown documents. See README for examples.
You may leave a comment below or discuss the post in the forum community.rstudio.com.